To use all functions of this page, please activate cookies in your browser.
my.chemeurope.com
With an accout for my.chemeurope.com you can always see everything at a glance – and you can configure your own website and individual newsletter.
- My watch list
- My saved searches
- My saved topics
- My newsletter
Phospholipase C
This article is about phosphoinositide phospholipase C (PI-PLC). For other types of phospholipase C enzymes, see Phospholipase C (disambiguation). Phosphoinositide phospholipase C (PLC) (EC 3.1.4.11) represents a family of eukaryotic intracellular enzymes that play an important role in signal transduction processes[1]. In general, this enzyme is denoted as phospholipase C, although three other families of phospholipase C enzymes have been identified in bacteria and in (eukaryotic) trypanosomes. Different phospholipase Cs participate in phosphatidylinositol bisphosphate (PIP2) metabolism and lipid signaling pathways in a calcium-dependent manner. At present, the superfamily consists of six sub-families comprising a total of 13 separate isoforms that differ in their mode of activation, catalytic regulation, cellular localization, membrane binding avidity, and tissue distribution. All are capable of catalyzing the hydrolysis of PIP2 into two important second messenger molecules, which go on to alter cell responses such as proliferation, differentiation, apoptosis, cytoskeleton remodeling, vesicular trafficking, ion channel conductance, endocrine function, and neurotransmission. Product highlight
Reaction and catalytic mechanismA phosphoinositide phospholipase C (EC 3.1.4.11) is an enzyme that catalyzes the chemical reaction
Thus, the two substrates of this enzyme are H2O and 1-phosphatidyl-1D-myo-inositol 4,5-bisphosphate (PIP2, phosphatidylinositol bisphosphate), whereas its two products are diacylglycerol and 1D-myo-inositol 1,4,5-trisphosphate (IP3, inositol triphosphate). All family members are capable of catalyzing the hydrolysis of PIP2, a phosphatidylinositol at the inner leaflet of the plasma membrane, generating two second messengers, inositol triphosphate (IP3) and diacylglycerol (DAG). PLCs catalyze the hydrolysis of PIP2 in two sequential steps. The first reaction is a phosphotransferase step that involves an intramolecular attack between the hydroxl group in the 2' position on the inositol ring and the phosphate group resulting in a cyclic IP3 intermediate. At this point, DAG is generated. However, in the second phosphodiesterase step, the cyclic intermediate is held within the active site long enough to be attacked by a molecule of water, resulting in a final acyclic IP3 product. It should be mentioned that bacterial forms of the enzyme, which contain only the catalytic lipase domain, produce cyclic intermediates exclusively, whereas the mammalian isoforms generate predominantly the acyclic product. However, it is possible to alter experimental conditions (e.g., temperature, pH) in vitro such that some mammalian isoforms will alter the degree to which they produce mixtures of cyclic/acyclic products along with DAG. RegulationThis catalytic process is tightly regulated by reversible phosphorylation and binding of regulatory proteins[2][3][4]. Cell locationPLCs perform their catalytic function at the plasma membrane where their substrate PIP2 is present. This membrane docking is mediated mostly by lipid-binding domains (e.g., PH domain, C2 domain) that display affinity for different phospholipid components of the plasma membrane. It is important to note that research has also discovered that, in addition to the plasma membrane, PLCs also exist within other sub-cellular regions such as the cytoplasm and nucleus of the cell. At present, it is unclear exactly what the definitive roles for these enzymes in these cellular compartments are, particularly the nucleus. FunctionPhospolipase C performs a catalytic mechanism, generating inositol triphosphate (IP3) and diacylglycerol (DAG). These molecules then go on to modulate the activity of downstream proteins important for cellular signaling. IP3 is soluble, and diffuses through the cytoplasm and interacts with IP3 receptors on the endoplasmic reticulum, causing the release of calcium and raising the level of intracellular calcium. Further reading: Function of calcium in humans DAG remains tethered to the inner leaflet of the plasma membrane due to its hydrophobic character, where it recruits protein kinase C (PKC), which becomes activated in conjunction with binding calcium ions. This results in a host of cellular responses through stimulation of calcium-sensitive proteins such as Calmodulin. Further reading: Function of protein kinase C Domain structureIn terms of domain organization, all family members possess homologous X and Y catalytic domains in the form of a distorted Triose Phosphate Isomerase (TIM) barrel with a highly-disordered, charged, and flexible intervening linker region. Likewise, all isoforms possess four EF hand domains, and a single C2 domain that flank the X and Y catalytic core. An N-terminal PH domain is present in every family except for the sperm-specific ζ isoform. SH2 (phosphotyrosine binding) and SH3 (proline-rich-binding) domains are found only in the γ form (specifically within the linker region), and only the ε form contains both guanine nucleotide exchange factor (GEF) and RA (Ras Associating) domains. The β subfamily is distinguished from the others by the presence of a long C-terminal extension immediately downstream of the C2 domain, which is required for activation by Gαq subunits, and which may also play a role in nuclear localization. Isozymes and activationThe Phospholipase C family consists of 13 isozymes split between six subfamilies, PLC-δ (1,3 & 4), -β(1-4), -γ(1,2), -ε, -ζ, and the recently-discovered -η(1,2) isoform. Depending on the specific subfamily in question, activation can be highly variable. Activation by either Gαq or Gβγ G-protein subunits (making it part of a G protein-coupled receptor signal transduction pathway) or by transmembrane receptors with intrinsic or associated tyrosine kinase activity has been reported. In addition, members of the Ras superfamily of small GTPases (namely the Ras and Rho subfamilies) have also been implicated. It should also be mentioned that all forms of Phospholipase C require calcium for activation, many of them possessing multiple calcium contact sites in the catalytic region. The only isoform that is known to be inactive at basal intracellular calcium levels is the δ subfamily of enzymes suggesting that they function as calcium amplifiers that become activated downstream of other PLC family members. PLC-δPLC-δ1 (85kDa) is activated by high calcium levels generated by other PLC family members, and therefore functions as a calcium amplifier within the cell. Binding of its substrate PIP2 to the N-terminal PH domain is highly specific and functions to promote activation of the catalytic core. In addition, this specificity helps tether the enzyme tightly to the plasma membrane in order to access substrate. While the catalytic core does possess a weak affinity for PIP2, the C2 domain has been shown to mediate calcium-dependent phospholipid binding as well. In this model, the PH and C2 domains operate in concert as a "tether and fix" apparatus necessary for processive catalysis by the enzyme. PLC-δ1 also possesses a classic leucine-rich Nuclear Export Signal (NES) in its EF hand motif, as well as a Nuclear Localization Signal within its linker region. These two elements combined allow PLC-δ1 to actively translocate in and out of the nucleus. However, its function in the nucleus remains obscure. Because it is the only family found expressed in lower eukaryotic organisms such as yeast and slime molds, it is considered the prototypical PLC isoform. The other family members more than likely evolved from PLC-δ as their domain architecture and mechanism of activation were expanded. Although a full crystal structure has not been obtained, high-resolution X-ray crystallography has yielded the molecular structure of the N-terminal PH domain complexed with IP3, as well as the remainder of the enzyme with the PH domain ablated. These structures have provided researchers with the necessary information to begin speculating about other family members. The widely-expressed PLC-δ1 isoform is the best-characterized phospholipase family member, as it was the first to have high-resolution X-ray crystal structures available for analysis. In terms of domain architecture, all of the enzymes are built upon a common PLC-δ backbone, wherein each family displays similarities, as well as obvious distinctions, that contribute to unique regulatory properties within the cell. PLC-βPLC-β(1-4) (120-155kDa) are activated by Gαq subunits through their C2 domain and long C-terminal extension. Gβγ subunits are known to activate the β2 and β3 isozymes only; however, this occurs through the PH domain. The exact mechanism still requires further investigation. The PH domain of β2 and β3 plays a duel role, much like PLC-δ1, by binding to the plasma membrane, as well being a site of interaction for the catalytic activator. However, PLC-β binds to the lipid surface independent of PIP2 with all isozymes preferring phosphoinositol-3-phosphate or neutral membranes. Recently, members of the Rho GTPase family (e.g., Rac1, Rac2, Rac3, and cdc42) have been implicated in their activation by binding to an alternate site on the N-terminal PH domain followed by subsequent recruitment to the plasma membrane. Like PLC-δ1, many PLC-β isoforms (in particular, PLC-β1) have been found to take up residence in the nuclear compartment. A basic amino acid region within the enzyme's long C-terminal tail appears to function as a Nuclear Localization Signal for import into the nucleus. PLC-β1 seems to play unspecified roles in cellular proliferation and differentiation. Other PLC families
Human proteins containing this domainPLCB1; PLCB2; PLCB3; PLCB4; PLCD1; PLCD3; PLCD4; PLCE1; PLCG1; PLCG2; PLCH1; PLCH2; PLCL1; PLCL2; PLCZ1; Systematic name and synonymsPhosphoinositide phospholipase C has several synonyms in common use, including triphosphoinositide phosphodiesterase, phosphoinositidase C, 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase, monophosphatidylinositol phosphodiesterase, phosphatidylinositol phospholipase C, PI-PLC, 1-phosphatidyl-D-myo-inositol-4,5-bisphosphate, and inositoltrisphosphohydrolase. Its systematic name is 1-phosphatidyl-1D-myo-inositol-4,5-bisphosphate inositoltrisphosphohydrolase. It belongs to the family of hydrolases, in specific, those acting on phosphoric diester bonds. References
See also
Additional images
Gene Ontology (GO) codes
Categories: EC 3.1.4 | Signal transduction | Peripheral membrane proteins | Enzymes of known structure |
||||||||||||||||||||||||||||||||||||||||
| This article is licensed under the GNU Free Documentation License. It uses material from the Wikipedia article "Phospholipase_C". A list of authors is available in Wikipedia. | ||||||||||||||||||||||||||||||||||||||||





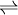 1D-myo-inositol 1,4,5-trisphosphate + diacylglycerol
1D-myo-inositol 1,4,5-trisphosphate + diacylglycerol


