To use all functions of this page, please activate cookies in your browser.
my.chemeurope.com
With an accout for my.chemeurope.com you can always see everything at a glance – and you can configure your own website and individual newsletter.
- My watch list
- My saved searches
- My saved topics
- My newsletter
Turn (biochemistry)A turn is an element of secondary structure in proteins. According to the most common definition, a turn is defined by the close approach of two Cα atoms (< 7 Å), when the corresponding residues are not involved in a regular secondary structure element such as an alpha helix or beta sheet. Product highlight
Types of turnsTurns are grouped by their hydrogen bonding and by their backbone dihedral angles. At the level of hydrogen bonds, the nomenclature is similar to that of helices.
Finally, an ω-loop is a catch-all term for a longer loop with no internal hydrogen bonding. Strictly speaking, a turn is defined by the close approach (< 7 Å) of Cα atoms and need not have a well-formed hydrogen bond. Thus, it is more correct to define a β-turn by the close approach of Cα atoms of residues separated by three peptide bonds, a γ-turn by the close approach of Cα atoms of residues separated by two peptide bonds, etc. In most cases, the H-bonding and Cα-distance definitions are equivalent. Within each hydrogen-bonding type, turns may be classified by their backbone dihedral angles. A turn can be converted into its inverse turn (also called its mirror-image turn) by changing the sign on all of its dihedral angles. (The inverse turn is not a true mirror image since the chirality of the Cα atoms is maintained.) Thus, the γ-turn has two forms, a classical form with (φ, ψ) dihedral angles of roughly Hairpins vs. diverging turnsA hairpin is a special case of a turn, in which the direction of the protein backbone reverses and the flanking secondary structure elements interact. For example, a β-hairpin connects two hydrogen-bonded, antiparallel β-strands. However, turns can cause less drastic changes in direction and may connect regular secondary structure elements that do not interact with each other. Such turns are called diverging turns. Role in protein foldingTwo hypotheses have been proposed for the role of turns in protein folding. In one view, turns play a critical role in folding by bringing together and fostering interactions between regular secondary structure elements. This view is supported by mutagenesis studies indicating a critical role for particular residues in the turns of some proteins. Also, nonnative isomers of X-Pro peptide bonds in turns can completely block the conformational folding of some proteins. In the opposing view, turns play a passive role in folding. This view is supported by the poor amino-acid conservation observed in most turns. Also, non-native isomers of many X-Pro peptide bonds in turns have little or no effect on folding. See alsoReferences
|
|||||||||||||||||||||||||||||
| This article is licensed under the GNU Free Documentation License. It uses material from the Wikipedia article "Turn_(biochemistry)". A list of authors is available in Wikipedia. | |||||||||||||||||||||||||||||





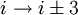 H-bonding).
H-bonding).
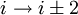 H-bonding).
H-bonding).
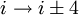 H-bonding).
H-bonding).
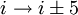 H-bonding).
H-bonding).
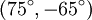 and an inverse form with dihedral angles
and an inverse form with dihedral angles 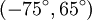 . At least eight forms of the β-turn have been identified, varying mainly in whether a cis isomer of a peptide bond is involved and on the dihedral angles of the central two residues. The classical and inverse β-turns are usually distinguished with a prime, e.g., type
. At least eight forms of the β-turn have been identified, varying mainly in whether a cis isomer of a peptide bond is involved and on the dihedral angles of the central two residues. The classical and inverse β-turns are usually distinguished with a prime, e.g., type 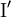 β-turns.
β-turns.


