DNA as a Template for Minerals
Copying DNA structures with silicates- a step toward ordered synthetic inorganic materials?
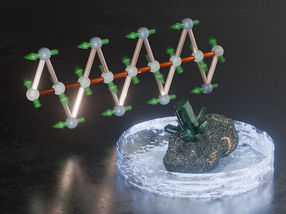
DNA carries the blueprint for all living things-and now it could also serve as the model for "dead" inorganic materials. However, the mechanism will be completely different from the usual "transcription" of the code by base pairs; instead, it is DNA's spatial structure that counts in this case. Japanese researchers have used the genetic molecule as a "scaffold" for the construction of tiny silicate rods and rings.
Looking at diatoms under a microscope is like looking into a bizarre microcosm: the skeletons of the tiny creatures consist of surprisingly precise silicate structures. Interest in such highly ordered inorganic nanostructures, which are considered to be potential catalysts or nanotech components, is strong; yet replication of these fossil structures has thus far seemed impossible. Whereas organic materials can be assembled into defined superstructures by the self-organization of smaller, tailored components, this is not possible for inorganic substances. The solution may be to use organic aggregates or large biomolecules as a starting point for the synthesis of inorganic structures.
A team led by Seiji Shinkai chose to use a plasmid DNA from bacteria as a matrix for the construction of tiny silicate structures. To pull this off, they needed to overcome two obstacles: The silicate precursors they used are negatively charged molecules, which will only stick to a positive matrix, and DNA also carries negative charges. In addition, the silicate precursor requires an organic solvent, while DNA is only soluble in water. The team was able to solve both problems in one elegant stroke: by attaching to the DNA a special ion, a hydrocarbon chain with positively charged groups at both its head and tail. The unusual thing is that the group at the chain's head is a guanidinium group, in which two nitrogen atoms "share" a positive charge so that they can simultaneously dock onto two oxygen atoms of a negatively charged phosphate group on the DNA backbone, a particularly favorable arrangement. The DNA phosphates' attraction to the guanidinium groups is thus much stronger than that for the second charge at the "tail" of the chain. This group thus remains free and gives the DNA a positive charge to attract and hold the silicate precursor. The hydrocarbon chains also impart the necessary solubility of the DNA in the organic solvent. Prepared in this manner, the plasmid DNA proved to be an ideal matrix for silicate formation. The silicate precursors attach to the DNA and then fuse together to form the silicate material. The DNA can subsequently be removed by heating. Plasmid DNA is usually twisted into a rodlike shape. However, enzymes can be used to relax it into a ring-shaped molecule, which makes it possible to obtain both rods and rings of silicate. Different DNA types may be used to create other nanosized "artificial fossils".
Other news from the department science
Most read news
More news from our other portals
See the theme worlds for related content
Topic world Synthesis
Chemical synthesis is at the heart of modern chemistry and enables the targeted production of molecules with specific properties. By combining starting materials in defined reaction conditions, chemists can create a wide range of compounds, from simple molecules to complex active ingredients.

Topic world Synthesis
Chemical synthesis is at the heart of modern chemistry and enables the targeted production of molecules with specific properties. By combining starting materials in defined reaction conditions, chemists can create a wide range of compounds, from simple molecules to complex active ingredients.