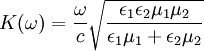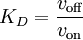To use all functions of this page, please activate cookies in your browser.
my.chemeurope.com
With an accout for my.chemeurope.com you can always see everything at a glance – and you can configure your own website and individual newsletter.
- My watch list
- My saved searches
- My saved topics
- My newsletter
Surface plasmon resonanceThe excitation of surface plasmons by light is denoted as a surface plasmon resonance (SPR) for planar surfaces or localized surface plasmon resonance (LSPR) for nanometer-sized metallic structures. This phenomenon is the basis of many standard tools for measuring adsorption of material onto planar metal (typically gold and silver) surfaces or onto the surface of metal nanoparticles. It is behind many color based biosensor applications and different lab-on-a-chip sensors.
Product highlight
ExplanationSurface plasmons, also known as surface plasmon polaritons, are surface electromagnetic waves that propagate parallel along a metal/dielectric (or metal/vacuum) interface. Since the wave is on the boundary of the metal and the external medium (air or water for example), these oscillations are very sensitive to any change of this boundary, such as the adsorption of molecules to the metal surface. To describe the existence and properties of surface plasmons, one can choose from various models (quantum theory, Drude model , etc.). The simplest way to approach the problem is to treat each material as a homogeneous continuum, described by a dielectric constant. With the terms of this description for electronic surface plasmons to exist, the real part of the dielectric constant of the metal must be negative and its magnitude must be greater than that of the dielectric. This condition is met in the IR-visible wavelength region for air/metal and water/metal interfaces (where the real dielectric constant of a metal is negative and that of air or water is positive). Realisation
In order to excite surface plasmons in a resonant manner, one can use an electron or light beam (visible and infrared are typical). The incoming beam has to match its impulse to that of the plasmon. In the case of p-polarized light (polarization occurs parallel to the surface), this is possible by passing the light through a block of glass to increase the wavenumber (and the impulse), and achieve the resonance at a given wavelength and angle. S-polarized (polarization occurs perpendicular to the surface) light cannot excite electronic surface plasmons. Electronic and magnetic surface plasmons obey the following dispersion relation: Typical metals that support surface plasmons are silver and gold, but metals such as copper, titanium, or chromium can also support surface plasmon generation. Using light to excite SP waves, there are two constructions which are well known. In the Otto setup, the light is shone on the wall of a glass block, typically a prism, and totally reflected. A thin metal (for example gold) film is positioned close enough, that the evanescent waves can interact with the plasma waves on the surface and excite the plasmons. In the Kretschmann configuration, the metal film is evaporated onto the glass block. The light is again illuminating from the glass, and an evanescent wave penetrates through the metal film. The plasmons are excited at the outer side of the film. This configuration is used in most practical applications. ApplicationsSurface plasmons have been used to enhance the surface sensitivity of several spectroscopic measurements including fluorescence, Raman scattering, and second harmonic generation. However, in their simplest form, SPR reflectivity measurements can be used to detect DNA or proteins by the changes in the local index of refraction upon adsorption of the target molecule to the metal surface. If the surface is patterned with different biopolymers, the technique is called Surface Plasmon Resonance Imaging (SPRI). For nanoparticles, localized surface plasmon oscillations can give rise to the intense colors of solutions of plasmon resonance nanoparticles and/or very intense scattering. Nanoparticles of noble metals exhibit strong ultraviolet-Visible absorption bands that are not present in the bulk metal. Shifts in this resonance due to changes in the local index of refraction upon adsorption of biopolymers to the nanoparticles can also be used to detect biopolymers such as DNA or proteins. Related complimentary techniques include plasmon waveguide resonance, QCM and Dual Polarisation Interferometry ExamplesLayer-by-layer self-assembly
One of the first common applications of surface plasmon resonance spectroscopy was the measurement of the thickness of adsorbed self-assembled nanofilms on gold substrates. An example is presented in 'figure' The resonance curves shift to higher angles as the thickness of the adsorbed film increases. This example is a 'static SPR' measurement. When higher speed observation is desired, one can select an angle right below the resonance point (the angle of minimum reflectance), and measure the reflectivity changes at that point. This is the so called 'dynamic SPR' measurement. The interpretation of the data assumes, that the structure of the film does not change significantly during the measurement. Binding constant determinationWhen the affinity of two ligands has to be determined, the binding constant must be determined. It is the equilibrium value for the product quotient. This value can also be found using the dynamical SPR parameters and, as in any chemical reaction, it is the association rate divided by the dissociation rate. For this, a so-called bait ligand is coated to the gold surface of the SPR crystal. Through a microflow system, a solution with the prey ligand can flow over the bait layer and bind. Binding will make the SPR signal change to an equilibrium. After some time, a solution without the prey is applied, and a new equilibrium will be reached. From these association ('on rate', von) and dissociation speeds ('off rate', voff), the binding constant can be calculated. The actual SPR signal can be explained by the electromagnetic 'coupling' of the incident light with the surface plasmon of the gold layer. This plasmon can be influenced by the layer just a few nanometer across the gold-solution interface i.e. the bait protein and possibly the prey protein. Binding makes the reflection angle change.;
Protein-Protein interaction
Experience shows that one RU represents the binding of approximately 1 pg/mm². In practice more thna 50 pg/mm² of analyte binding is needed (100 RU of binding should be adequate). SPR provides a good estimation of kon and koff if mass-transport and re-binding are insignificant, otherwise the apparent kon and koff values would be only lower estimates. There are two major ways to measure KD:
A useful rule of thumb is that an interaction should reach 99% of the equilibrium level within 4.6/koff s. There are three main types of coupling chemistry, which utilize, respectively, amine (e.g. lysine), thiol (cysteine), or aldehyde (carbohydrate; if the protein is glycosylated, i.e., glycoprotein. All covalent coupling methods utilize free carboxymethyl groups on the sensor chip surface. As controls use:
Perform all measurements in physiological temperature. Ideally analyte concentration should be varied over four orders of magnitude, from 0.01KD to 100KD. You may use Biaevaluation, Scrubber, or any other software to analyze your data and prepare it for presentation. All buffers should be filtered through 0.2 micrometer filters and degassed at room temperature. Data processing for measuring binding affinity1) Zeroing: zero on the y-axis for start time of the injection; 2) Reference subtraction: subtract-out data from the reference spots; 3) Replication overlays and double Referencing; 4) Globally fitting data: numerically fit the data to 1:1 Langmuir interaction model. Possible artifacts while measuring binding affinity1) Spikes in the sensogram in the response to elevated temperatures that can occur due to outgassing of the buffer; 2) excluded-volume effects; 3) Protein immobilization affects affinity; 4) the injected analyte is not monomeric (use analytical ultracentrifugation to assure monomeric state, also use gel filtration), aggregation; 5) avidity effects; 6) both analyte and ligand should be chemically and conformationally pure; 7) instrument is not clean and should be primed; 8) baseline drift (use covalent immobilization); 9) check for non-specific binding; 10) surface heterogeneity (look at the shape of the 'SPR dip'); 11) surface instability (baseline drift); 12) mass-transport effects, check whether kon and koff dependence on flow rate, use high flow rates (100 µL/min) and low levels of immobilized ligand; 13) one cannot measure association or dissociation rates which are faster than the machine sampling frequency (usually 1 Hz); 14) If your concentrations are wrong then your affinity results would be wrong [concentration of correctly folded protein]; 15) If your protein Lysine content is high you might consider conjugating the protein by a method other than amine chemistry [biotin is recommended]; 16) degradarion of the protein on the chip during long measurements (several hours); 17) Biphasic sensogram is bad; lower concentration to achieve a monotonic curve of association and dissociation. Validation of measuring binding affinity1) Isothermal titration calorimetry; 2) Nuclear magnetic resonance; 3) using different machine (Biacore vs. ProteOn); 4) co-immunoprecipitation (less quantitative) Tips for achieving a good measurement of binding affinity1) To avoid using too much protein inject a small amount of sample only for a short time [even if you don't reach a steady-state]. That will help avoiding aggregation effects; 2) If the two binding partners are protein and short peptide, conjugate the peptide to the surface. Otherwise, several peptides would bind a single protein in mixed specific and non-specific binding; 3) Add 1% alignate top the buffer to reduce non-specific association; 4) protein must be fresh (i.e., not frozen), after gel-filtration, and kept on ice; 5) use as a reference measurement "empty but activated" surface; 6) chi-squared should be lower than 10% of Rmax. Magnetic Plasmon ResonanceRecently, there has been an interest in magnetic surface plasmons. These require materials with large negative magnetic permeability, a property that has only recently been made available with the construction of metamaterials. References
See also
Categories: Nanotechnology | Spectroscopy | Biochemistry methods |
|||
| This article is licensed under the GNU Free Documentation License. It uses material from the Wikipedia article "Surface_plasmon_resonance". A list of authors is available in Wikipedia. |