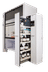Pond-dwelling powerhouse's genome points to its biofuel potential
Advertisement
Duckweed is a tiny floating plant that's been known to drive people daffy. It's one of the smallest and fastest-growing flowering plants that often becomes a hard-to-control weed in ponds and small lakes. But it's also been exploited to clean contaminated water and as a source to produce pharmaceuticals. Now, the genome of Greater Duckweed (Spirodela polyrhiza) has given this miniscule plant's potential as a biofuel source a big boost. In a paper published in the journal Nature Communications, researchers from Rutgers University, the Department of Energy Joint Genome Institute and several other facilities detailed the complete genome of S. polyrhiza and analyzed it in comparison to several other plants, including rice and tomatoes.

Duckweed is a relatively simple plant with fronds that float on the surface of the water and roots that extend into the water. In the flask on the left, you can see the dormant phase, turions, that have dropped to the bottom.
Wenquin Wang

Duckweed, a small, common plant that grows in ponds and stagnant waters, is an ideal candidate as a biofuel raw material.
Texx Smith, via flickr
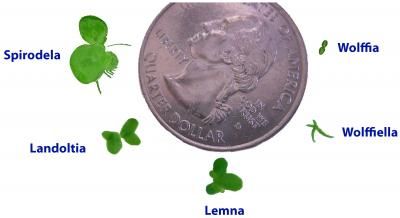
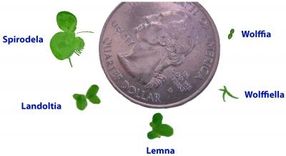
Spirodela is one of the smallest plants in the world. Here it is displayed with other comparable plants.
Wang W, Kerstetter RA, Michael TP. Evolution of Genome Size in Duckweeds (Lemnaceae). Journal of Botany 2011, 1-9 (2011).


Simple and primitive, a duckweed plant consists of a single small kidney-shaped leaf about the size of a pencil-top eraser that floats on the surface of the water with a few thin roots underwater. It grows in almost all geographic areas, at nearly any altitude. Although it's a flowering plant, it only rarely forms small indistinct flowers on the underside of its floating leaves. Most of the time, it reproduces by budding off small leaves that are clones of the parent leaf. It often forms thick mats on the edges of ponds, quiet inlets of lakes and in marshes. It's among the fastest growing plants, able to double its population in a couple of days under ideal conditions.
These and other properties make it an ideal candidate as a biofuel feedstock – a raw source for biofuel production. For example, unlike plants on land, duckweeds don't need to hold themselves upright or transport water from distant roots to their leaves, so they're a relatively soft and pliable plant, containing tiny amounts of woody material such as lignin and cellulose. Removing these woody materials from feedstock has been a major challenge in biofuel production. Also, although they are small enough to grow in many environments, unlike biofuel-producing microbes, duckweed plants are large enough to harvest easily.
S. polyrhiza turns out to have one of the smallest known plant genomes, at about 158 million base pairs and fewer than 20,000 protein-encoding genes. That's 27 percent fewer than Arabidopsis thaliana – which, until recently, was believed to be the smallest plant genome – and nearly half as many as rice plants.
"The most surprising find was insight into the molecular basis for genes involved in maturation – a forever-young lifestyle," said senior author Joachim Messing, director of the Waksman Institute of Microbiology at Rutgers University.
S. polyrhiza leaves resemble cotyledons, embryonic leaves inside plant seeds that become the first leaves after germination. But where other plants develop other kinds of leaves as they mature, S. polyrhiza's never progresses and continuously produces cotyledon leaves. This prolonging of juvenile traits is called "neoteny." S. polyrhiza had fewer genes to promote and more genes to repress the switch from juvenile to mature growth.
"Because of the reduction in neoteny, there is an arrest in development and differentiation of organs. So this arrest allowed us to uncover regulatory networks that are required for differentiation and development," Messing said.
Also intriguing to the research team were which genes were preserved over time and which were not. Many of the genes responsible for cellulose and lignin production in land dwelling plants were missing, and there were fewer copies of those that were present. Genes for another compound related to cell walls called "expansins" which are involved with cell wall and root growth were also reduced.
Genes for starch production, on the other hand, were retained and are probably used for creating starch-filled turions, specialized buds produced by aquatic plants for overwintering, enabling them sink to the bottom of ponds and revive in warmer weather. Moreover, despite the reduced number of total genes, S. polyrhiza has more copies of genes for enzymes involved in nitrogen absorption and metabolism than in other plants. This is probably linked to the plant's ability to utilize excess nitrogen in contaminated waters.
A thorough understanding of the genome and cellular mechanisms of S. polyrhiza could greatly enhance current efforts to recruit duckweed as a biofuel source. Messing estimates that duckweed will be a viable biofuel source within the next five years and points to Ceres Energy Group in New Jersey, which is already producing electricity from duckweed. Understanding which genes produce which traits will allow researchers to create new varieties of duckweed with enhanced biofuel traits, such as increased reduction of cellulose or increased starch or even higher lipid production. Starch can be directly used as a biofuel source and it can be converted to ethanol, the way corn is currently converted to ethanol fuel, but oils would have greater energy than ethanol.
"Classical breeding or genetics does not apply here because of its clonal propagation and rare flowering, but these organisms can be transformed with DNA," Messing said. "Therefore, new variants can be created with modified pathways for industrial applications. These variants would be an enhancement over what can be done now."
This genome was sequenced as part of a DOE Office of Science JGI Community Science Program (CSP) project (formerly the Community Sequencing Program). It exemplifies the collaborative approach and innovative projects that the CSP enables among researchers. Messing pointed to the study's advances over previous research.
"The sequencing of this genome opens new frontiers in the molecular biology of aquatic plants," said Messing. "This publication represents the single largest advance in this field and a new milestone in plant molecular biology and evolution, as previous studies were either classical botany or biochemistry of photosynthesis. The placement of the Spirodela genome as a basal monocot species will serve as a new reference for all flowering plants."